plotting example: plot_2d_boxes¶
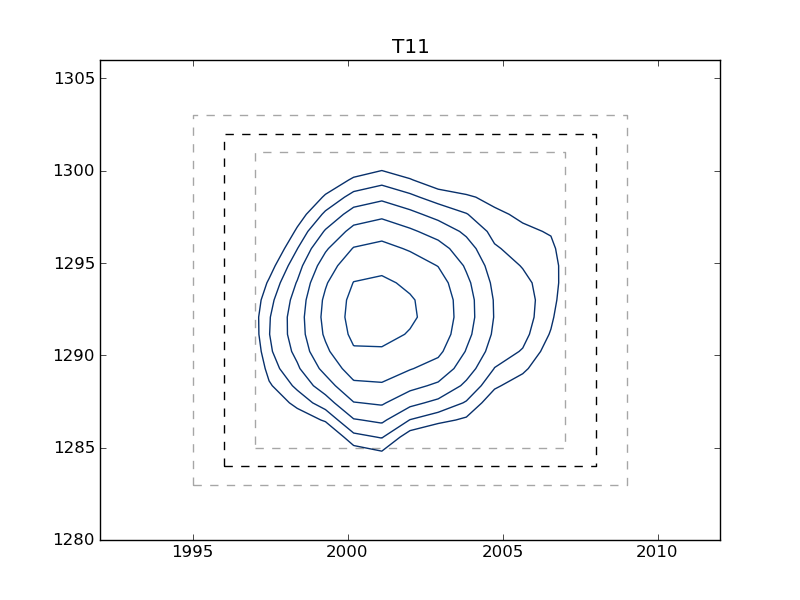
This example shows how to use nmrglue and matplotlib to create figures for examining data or publication. In this example the box limits used in integration example: integrate_2d are graphically examined. A contour plot of each peak is plotted with the box limits indicated by the dark dashed line. To check peak assignments see plotting example: plot_2d_assignments.
The data used in this example is available for download.
#! /usr/bin/env python
# Create a contour plots of each peak defined in limits.in file
import nmrglue as ng
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm
# plot parameters
xpad = 5 # padding around peak box on x-axis
ypad = 5 # padding around peak box on y-axis
cmap = matplotlib.cm.Blues_r # contour map (colors to use for contours)
contour_start = 30000 # contour level start value
contour_num = 20 # number of contour levels
contour_factor = 1.20 # scaling factor between contour levels
# calculate contour levels
cl = contour_start * contour_factor ** np.arange(contour_num)
# read in the data from a NMRPipe file
dic, data = ng.pipe.read("nmrpipe_2d/test.ft2")
# read in the integration limits
peak_list = np.recfromtxt("limits.in")
# loop over the peaks
for name, x0, y0, x1, y1 in peak_list:
if x0 > x1:
x0, x1 = x1, x0
if y0 > y1:
y0, y1 = y1, y0
# slice the data around the peak
slice = data[y0 - ypad:y1 + 1 + ypad, x0 - xpad:x1 + 1 + xpad]
# create the figure
fig = plt.figure()
ax = fig.add_subplot(111)
# plot the contours
print "Plotting:", name
etup = (x0 - xpad + 1, x1 + xpad - 1, y0 - ypad + 1, y1 + ypad - 1)
ax.contour(slice, cl, cmap=cmap, extent=etup)
# draw a box around the peak
ax.plot([x0, x1, x1, x0, x0], [y0, y0, y1, y1, y0], 'k--')
# draw light boxes at +/- one point
ax.plot([x0 - 1, x1 + 1, x1 + 1, x0 - 1, x0 - 1],
[y0 - 1, y0 - 1, y1 + 1, y1 + 1, y0 - 1], 'k--', alpha=0.35)
ax.plot([x0 + 1, x1 - 1, x1 - 1, x0 + 1, x0 + 1],
[y0 + 1, y0 + 1, y1 - 1, y1 - 1, y0 + 1], 'k--',alpha=0.35)
# set the title
ax.set_title(name)
# save the figure
fig.savefig(name + ".png")
del(fig)
#Peak X0 Y0 X1 Y1
# Peak defines 15N resonance in 2D NCO spectra.
# Limits are in term of points from 0 to length-1.
# These can determined from nmrDraw by subtracting 1 from the X and Y
# values reported.
#Peak X0 Y0 X1 Y1
T49 1992 1334 2003 1316
T11 1996 1302 2008 1284
# comments can appear anywhere in this file just start the line with #
G14 2032 1314 2044 1293
E15 2077 1025 2087 1004
W43 2008 952 2019 933
Sample Figure
[T11.png]